Network Layers¶
Layer definition¶
Common set of parameters for any kind of layer.
Option [default value] |
Description |
|---|---|
|
Name of the section(s) for the input layer(s). Comma separated |
|
Type of the layer. Can be any of the type described below |
|
Layer model to use |
|
Layer data type to use. Please note that some layers may not support every data type. |
|
Name of the configuration section for layer |
Weight fillers¶
Fillers to initialize weights and biases in the different type of layer.
Usage example:
[conv1]
...
WeightsFiller=NormalFiller
WeightsFiller.Mean=0.0
WeightsFiller.StdDev=0.05
...
The initial weights distribution for each layer can be checked in the weights_init folder, with an example shown in figure [fig:weightsInitDistrib].
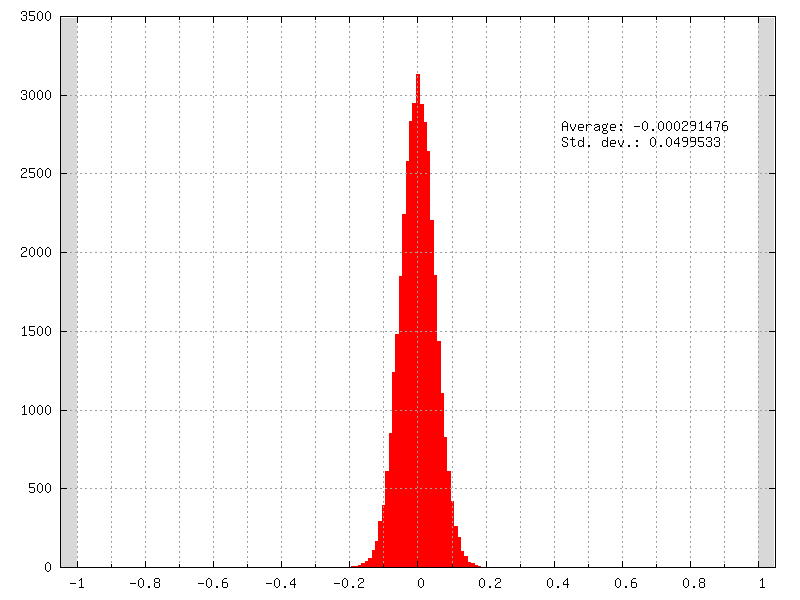
Initial weights distribution of a layer using a normal distribution
(NormalFiller) with a 0 mean and a 0.05 standard deviation.¶
ConstantFiller¶
Fill with a constant value.
Option |
Description |
|---|---|
FillerName |
Value for the filling |
HeFiller¶
Fill with an normal distribution with normalized variance taking into account the rectifier nonlinearity [HZRS15]. This filler is sometimes referred as MSRA filler.
Option [default value] |
Description |
|---|---|
FillerName |
Normalization, can be |
FillerName |
Scaling factor |
Use a normal distribution with standard deviation \(\sqrt{\frac{2.0}{n}}\).
\(n\) = \(fan\text{-}in\) with
FanIn, resulting in \(Var(W)=\frac{2}{fan\text{-}in}\)\(n\) = \(\frac{(fan\text{-}in + fan\text{-}out)}{2}\) with
Average, resulting in \(Var(W)=\frac{4}{fan\text{-}in + fan\text{-}out}\)\(n\) = \(fan\text{-}out\) with
FanOut, resulting in \(Var(W)=\frac{2}{fan\text{-}out}\)
NormalFiller¶
Fill with a normal distribution.
Option [default value] |
Description |
|---|---|
FillerName |
Mean value of the distribution |
FillerName |
Standard deviation of the distribution |
UniformFiller¶
Fill with an uniform distribution.
Option [default value] |
Description |
|---|---|
FillerName |
Min. value |
FillerName |
Max. value |
XavierFiller¶
Fill with an uniform distribution with normalized variance [GB10].
Option [default value] |
Description |
|---|---|
FillerName |
Normalization, can be |
FillerName |
Distribution, can be |
FillerName |
Scaling factor |
Use an uniform distribution with interval \([-scale,scale]\), with \(scale = \sqrt{\frac{3.0}{n}}\).
\(n\) = \(fan\text{-}in\) with
FanIn, resulting in \(Var(W)=\frac{1}{fan\text{-}in}\)\(n\) = \(\frac{(fan\text{-}in + fan\text{-}out)}{2}\) with
Average, resulting in \(Var(W)=\frac{2}{fan\text{-}in + fan\text{-}out}\)\(n\) = \(fan\text{-}out\) with
FanOut, resulting in \(Var(W)=\frac{1}{fan\text{-}out}\)
Weight solvers¶
SGDSolver_Frame¶
SGD Solver for Frame models.
Option [default value] |
Description |
|---|---|
SolverName |
Learning rate |
SolverName |
Momentum |
SolverName |
Decay |
SolverName |
Learning rate decay policy. Can be any of |
SolverName |
Learning rate step size (in number of stimuli) |
SolverName |
Learning rate decay |
SolverName |
If true, clamp the weights and bias between -1 and 1 |
SolverName |
Polynomial learning rule power parameter |
SolverName |
Polynomial learning rule maximum number of iterations |
The learning rate decay policies are the following:
StepDecay: every SolverName.LearningRateStepSizestimuli, the learning rate is reduced by a factor SolverName.LearningRateDecay;ExponentialDecay: the learning rate is \(\alpha = \alpha_{0}\exp(-k t)\), with \(\alpha_{0}\) the initial learning rate SolverName.LearningRate, \(k\) the rate decay SolverName.LearningRateDecayand \(t\) the step number (one step every SolverName.LearningRateStepSizestimuli);InvTDecay: the learning rate is \(\alpha = \alpha_{0} / (1 + k t)\), with \(\alpha_{0}\) the initial learning rate SolverName.LearningRate, \(k\) the rate decay SolverName.LearningRateDecayand \(t\) the step number (one step every SolverName.LearningRateStepSizestimuli).InvDecay: the learning rate is \(\alpha = \alpha_{0} * (1 + k t)^{-n}\), with \(\alpha_{0}\) the initial learning rate SolverName.LearningRate, \(k\) the rate decay SolverName.LearningRateDecay, \(t\) the current iteration and \(n\) the power parameter SolverName.PowerPolyDecay: the learning rate is \(\alpha = \alpha_{0} * (1 - \frac{k}{t})^n\), with \(\alpha_{0}\) the initial learning rate SolverName.LearningRate, \(k\) the current iteration, \(t\) the maximum number of iteration SolverName.MaxIterationsand \(n\) the power parameter SolverName.Power
SGDSolver_Frame_CUDA¶
SGD Solver for Frame_CUDA models.
Option [default value] |
Description |
|---|---|
SolverName |
Learning rate |
SolverName |
Momentum |
SolverName |
Decay |
SolverName |
Learning rate decay policy. Can be any of |
SolverName |
Learning rate step size (in number of stimuli) |
SolverName |
Learning rate decay |
SolverName |
If true, clamp the weights and bias between -1 and 1 |
The learning rate decay policies are identical to the ones in the
SGDSolver_Frame solver.
AdamSolver_Frame¶
Adam Solver for Frame models [KB14].
Option [default value] |
Description |
|---|---|
SolverName |
Learning rate (stepsize) |
SolverName |
Exponential decay rate of these moving average of the first moment |
SolverName |
Exponential decay rate of these moving average of the second moment |
SolverName |
Epsilon |
AdamSolver_Frame_CUDA¶
Adam Solver for Frame_CUDA models [KB14].
Option [default value] |
Description |
|---|---|
SolverName |
Learning rate (stepsize) |
SolverName |
Exponential decay rate of these moving average of the first moment |
SolverName |
Exponential decay rate of these moving average of the second moment |
SolverName |
Epsilon |
Activation functions¶
Activation function to be used at the output of layers.
Usage example:
[conv1]
...
ActivationFunction=Rectifier
ActivationFunction.LeakSlope=0.01
ActivationFunction.Clipping=20
...
Logistic¶
Logistic activation function.
LogisticWithLoss¶
Logistic with loss activation function.
Rectifier¶
Rectifier or ReLU activation function.
Option [default value] |
Description |
|---|---|
|
Leak slope for negative inputs |
|
Clipping value for positive outputs |
Saturation¶
Saturation activation function.
Softplus¶
Softplus activation function.
Tanh¶
Tanh activation function.
Computes \(y = tanh(\alpha x)\).
Option [default value] |
Description |
|---|---|
|
\(\alpha\) parameter |
TanhLeCun¶
Tanh activation function with an \(\alpha\) parameter of \(1.7159 \times (2.0/3.0)\).
Anchor¶
Anchor layer for Faster R-CNN or Single Shot Detector.
Option [default value] |
Description |
|---|---|
|
This layer takes one or two inputs. The total number of input channels must be |
|
Anchors definition. For each anchor, there must be two space-separated values: the root area and the aspect ratio. |
|
Number of classes per anchor. Must be 1 (if the scores input uses logistic regression) or 2 (if the scores input is a two-class softmax layer) |
|
Reference width use to scale anchors coordinate. |
|
Reference height use to scale anchors coordinate. |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Assign a positive label for anchors whose IoU overlap is higher than |
|
all Frame |
Assign a negative label for non-positive anchors whose IoU overlap is lower than |
|
all Frame |
Balancing parameter \(\lambda\) |
|
all Frame |
Number of random positive samples for the loss computation |
|
all Frame |
Number of random negative samples for the loss computation |
Usage example:
; RPN network: cls layer
[scores]
Input=...
Type=Conv
KernelWidth=1
KernelHeight=1
; 18 channels for 9 anchors
NbOutputs=18
...
[scores.softmax]
Input=scores
Type=Softmax
NbOutputs=[scores]NbOutputs
WithLoss=1
; RPN network: coordinates layer
[coordinates]
Input=...
Type=Conv
KernelWidth=1
KernelHeight=1
; 36 channels for 4 coordinates x 9 anchors
NbOutputs=36
...
; RPN network: anchors
[anchors]
Input=scores.softmax,coordinates
Type=Anchor
ScoresCls=2 ; using a two-class softmax for the scores
Anchor[0]=32 1.0
Anchor[1]=48 1.0
Anchor[2]=64 1.0
Anchor[3]=80 1.0
Anchor[4]=96 1.0
Anchor[5]=112 1.0
Anchor[6]=128 1.0
Anchor[7]=144 1.0
Anchor[8]=160 1.0
ConfigSection=anchors.config
[anchors.config]
PositiveIoU=0.7
NegativeIoU=0.3
LossLambda=1.0
Outputs remapping¶
Outputs remapping allows to convert scores and coordinates output
feature maps layout from another ordering that the one used in the N2D2
Anchor layer, during weights import/export.
For example, lets consider that the imported weights corresponds to the following output feature maps ordering:
0 anchor[0].y
1 anchor[0].x
2 anchor[0].h
3 anchor[0].w
4 anchor[1].y
5 anchor[1].x
6 anchor[1].h
7 anchor[1].w
8 anchor[2].y
9 anchor[2].x
10 anchor[2].h
11 anchor[2].w
The output feature maps ordering required by the Anchor layer is:
0 anchor[0].x
1 anchor[1].x
2 anchor[2].x
3 anchor[0].y
4 anchor[1].y
5 anchor[2].y
6 anchor[0].w
7 anchor[1].w
8 anchor[2].w
9 anchor[0].h
10 anchor[1].h
11 anchor[2].h
The feature maps ordering can be changed during weights import/export:
; RPN network: coordinates layer
[coordinates]
Input=...
Type=Conv
KernelWidth=1
KernelHeight=1
; 36 channels for 4 coordinates x 9 anchors
NbOutputs=36
...
ConfigSection=coordinates.config
[coordinates.config]
WeightsExportFormat=HWCO ; Weights format used by TensorFlow
OutputsRemap=1:4,0:4,3:4,2:4
BatchNorm¶
Batch Normalization layer [IS15].
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Activation function. Can be any of |
|
Share the scales with an other layer |
|
Share the biases with an other layer |
|
Share the means with an other layer |
|
Share the variances with an other layer |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Any solver parameters |
|
all Frame |
Scale solver parameters, take precedence over the |
|
all Frame |
Bias solver parameters, take precedence over the |
|
all Frame |
Epsilon value used in the batch normalization formula. If 0.0, automatically choose the minimum possible value. |
|
all Frame |
MovingAverageMomentum: used for the moving average of batch-wise means and standard deviations during training. The closer to 1.0, the more it will depend on the last batch. |
Conv¶
Convolutional layer.
Option [default value] |
Description |
|---|---|
|
Width of the kernels |
|
Height of the kernels |
|
Depth of the kernels (implies 3D kernels) |
|
Kernels size (implies 2D square kernels) |
|
List of space-separated dimensions for N-D kernels |
|
Number of output channels |
|
X-axis subsampling factor of the output feature maps |
|
Y-axis subsampling factor of the output feature maps |
|
Z-axis subsampling factor of the output feature maps |
|
Subsampling factor of the output feature maps |
|
List of space-separated subsampling dimensions for N-D kernels |
|
X-axis stride of the kernels |
|
Y-axis stride of the kernels |
|
Z-axis stride of the kernels |
|
Stride of the kernels |
|
List of space-separated stride dimensions for N-D kernels |
|
X-axis input padding |
|
Y-axis input padding |
|
Z-axis input padding |
|
Input padding |
|
List of space-separated padding dimensions for N-D kernels |
|
X-axis dilation of the kernels |
|
Y-axis dilation of the kernels |
|
Z-axis dilation of the kernels |
|
Dilation of the kernels |
|
List of space-separated dilation dimensions for N-D kernels |
|
Activation function. Can be any of |
|
Weights initial values filler |
|
Biases initial values filler |
|
Mapping: number of groups (mutually exclusive with all other Mapping.* options) |
|
Mapping: number of channels per group (mutually exclusive with all other Mapping.* options) |
|
Mapping canvas pattern default width |
|
Mapping canvas pattern default height |
|
Mapping canvas pattern default size (mutually
exclusive with |
|
Mapping canvas default X-axis step |
|
Mapping canvas default Y-axis step |
|
Mapping canvas default step (mutually exclusive
with``Mapping.StrideX`` and |
|
Mapping canvas default X-axis offset |
|
Mapping canvas default Y-axis offset |
|
Mapping canvas default offset (mutually exclusive
with |
|
Mapping canvas pattern default number of iterations (0 means no limit) |
|
Mapping canvas pattern default width for
input layer |
|
Mapping canvas pattern default height for
input layer |
|
Mapping canvas pattern default size for
input layer |
|
Mapping canvas default X-axis step for
input layer |
|
Mapping canvas default Y-axis step for
input layer |
|
Mapping canvas default step for input layer |
|
Mapping canvas default X-axis offset for
input layer |
|
Mapping canvas default Y-axis offset for
input layer |
|
Mapping canvas default offset for input
layer |
|
Mapping canvas pattern default number of
iterations for input layer |
|
Share the weights with an other layer |
|
Share the biases with an other layer |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
If true, don’t use bias |
|
all Frame |
Any solver parameters |
|
all Frame |
Weights solver parameters, take precedence over the |
|
all Frame |
Bias solver parameters, take precedence over the |
|
all Frame |
Weights import/export format. Can be |
|
all Frame |
If true, import/export flipped kernels |
Configuration parameters (Spike models)¶
Experimental option (implementation may be wrong or susceptible to change)
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Spike |
Synaptic incoming delay \(w_{delay}\) |
|
|
Threshold of the neuron \(I_{thres}\) |
|
|
If true, the threshold is also applied to the absolute value of negative values (generating negative spikes) |
|
|
Neural leak time constant \(\tau_{leak}\) (if 0, no leak) |
|
|
Neural refractory period \(T_{refrac}\) |
|
|
Relative initial synaptic weight \(w_{init}\) |
|
|
Mean minimum synaptic weight \(w_{min}\) |
|
|
Mean maximum synaptic weight \(w_{max}\) |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
Intrinsic SET switching probability \(P_{SET}\) (upon receiving a SET programming pulse). Assuming uniform statistical distribution (not well supported by experiments on RRAM) |
|
|
Intrinsic RESET switching probability \(P_{RESET}\) (upon receiving a RESET programming pulse). Assuming uniform statistical distribution (not well supported by experiments on RRAM) |
|
|
Synaptic redundancy (number of RRAM device per synapse) |
|
|
Bipolar weights |
|
|
Bipolar integration |
|
|
Extrinsic STDP LTP probability (cumulative with intrinsic SET switching probability \(P_{SET}\)) |
|
|
Extrinsic STDP LTD probability (cumulative with intrinsic RESET switching probability \(P_{RESET}\)) |
|
|
STDP LTP time window \(T_{LTP}\) |
|
|
Neural lateral inhibition period \(T_{inhibit}\) |
|
|
If false, STDP is disabled (no synaptic weight change) |
|
|
If true, reset the integration to 0 during the refractory period |
|
|
If false, the analog value of the devices is integrated, instead of their binary value |
Deconv¶
Deconvolution layer.
Option [default value] |
Description |
|---|---|
|
Width of the kernels |
|
Height of the kernels |
|
Depth of the kernels (implies 3D kernels) |
|
Kernels size (implies 2D square kernels) |
|
List of space-separated dimensions for N-D kernels |
|
Number of output channels |
|
X-axis subsampling factor of the output feature maps |
|
Y-axis subsampling factor of the output feature maps |
|
Z-axis subsampling factor of the output feature maps |
|
Subsampling factor of the output feature maps |
|
List of space-separated subsampling dimensions for N-D kernels |
|
X-axis stride of the kernels |
|
Y-axis stride of the kernels |
|
Z-axis stride of the kernels |
|
Stride of the kernels |
|
List of space-separated stride dimensions for N-D kernels |
|
X-axis input padding |
|
Y-axis input padding |
|
Z-axis input padding |
|
Input padding |
|
List of space-separated padding dimensions for N-D kernels |
|
X-axis dilation of the kernels |
|
Y-axis dilation of the kernels |
|
Z-axis dilation of the kernels |
|
Dilation of the kernels |
|
List of space-separated dilation dimensions for N-D kernels |
|
Activation function. Can be any of |
|
Weights initial values filler |
|
Biases initial values filler |
|
Mapping: number of groups (mutually exclusive with all other Mapping.* options) |
|
Mapping: number of channels per group (mutually exclusive with all other Mapping.* options) |
|
Mapping canvas pattern default width |
|
Mapping canvas pattern default height |
|
Mapping canvas pattern default size (mutually
exclusive with |
|
Mapping canvas default X-axis step |
|
Mapping canvas default Y-axis step |
|
Mapping canvas default step (mutually exclusive
with``Mapping.StrideX`` and |
|
Mapping canvas default X-axis offset |
|
Mapping canvas default Y-axis offset |
|
Mapping canvas default offset (mutually exclusive
with |
|
Mapping canvas pattern default number of iterations (0 means no limit) |
|
Mapping canvas pattern default width for
input layer |
|
Mapping canvas pattern default height for
input layer |
|
Mapping canvas pattern default size for
input layer |
|
Mapping canvas default X-axis step for
input layer |
|
Mapping canvas default Y-axis step for
input layer |
|
Mapping canvas default step for input layer |
|
Mapping canvas default X-axis offset for
input layer |
|
Mapping canvas default Y-axis offset for
input layer |
|
Mapping canvas default offset for input
layer |
|
Mapping canvas pattern default number of
iterations for input layer |
|
Share the weights with an other layer |
|
Share the biases with an other layer |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
If true, don’t use bias |
|
all Frame |
If true, enable backpropagation |
|
all Frame |
Any solver parameters |
|
all Frame |
Weights solver parameters, take precedence over the |
|
all Frame |
Bias solver parameters, take precedence over the |
|
all Frame |
Weights import/export format. Can be |
|
all Frame |
If true, import/export flipped kernels |
Dropout¶
Dropout layer [SHK+12].
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
The probability with which the value from input would be dropped |
ElemWise¶
Element-wise operation layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Type of operation ( |
|
Weights for the |
|
Shifts for the |
|
Activation function. Can be any of |
Given \(N\) input tensors \(T_{i}\), performs the following operation:
Sum operation¶
\(T_{out} = \sum_{1}^{N}(w_{i} T_{i} + s_{i})\)
AbsSum operation¶
\(T_{out} = \sum_{1}^{N}(w_{i} |T_{i}|)\)
EuclideanSum operation¶
\(T_{out} = \sqrt{\sum_{1}^{N}\left(w_{i} T_{i} + s_{i}\right)^{2}}\)
Prod operation¶
\(T_{out} = \prod_{1}^{N}(T_{i})\)
Max operation¶
\(T_{out} = MAX_{1}^{N}(T_{i})\)
Examples¶
Sum of two inputs (\(T_{out} = T_{1} + T_{2}\)):
[elemwise_sum]
Input=layer1,layer2
Type=ElemWise
NbOutputs=[layer1]NbOutputs
Operation=Sum
Weighted sum of two inputs, by a factor 0.5 for layer1 and 1.0 for
layer2 (\(T_{out} = 0.5 \times T_{1} + 1.0 \times T_{2}\)):
[elemwise_weighted_sum]
Input=layer1,layer2
Type=ElemWise
NbOutputs=[layer1]NbOutputs
Operation=Sum
Weights=0.5 1.0
Single input scaling by a factor 0.5 and shifted by 0.1 (\(T_{out} = 0.5 \times T_{1}\) + 0.1):
[elemwise_scale]
Input=layer1
Type=ElemWise
NbOutputs=[layer1]NbOutputs
Operation=Sum
Weights=0.5
Shifts=0.1
Absolute value of an input (\(T_{out} = |T_{1}|\)):
[elemwise_abs]
Input=layer1
Type=ElemWise
NbOutputs=[layer1]NbOutputs
Operation=Abs
FMP¶
Fractional max pooling layer [Gra14].
Option [default value] |
Description |
|---|---|
|
Number of output channels |
|
Scaling ratio. The output size is \(round\left(\frac{\text{input size}}{\text{scaling ratio}}\right)\). |
|
Activation function. Can be any of |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
If true, use overlapping regions, else use disjoint regions |
|
all Frame |
If true, use pseudorandom sequences, else use random sequences |
Fc¶
Fully connected layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Weights initial values filler |
[ |
|
|
Biases initial values filler |
[ |
|
|
Activation function. Can be any of |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
If true, don’t use bias |
|
all Frame |
If true, enable backpropagation |
|
all Frame |
Any solver parameters |
|
all Frame |
Weights solver parameters, take precedence over the |
|
all Frame |
Bias solver parameters, take precedence over the |
|
|
If below 1.0, fraction of synapses that are disabled with drop connect |
Configuration parameters (Spike models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Spike |
Synaptic incoming delay \(w_{delay}\) |
|
|
Threshold of the neuron \(I_{thres}\) |
|
|
If true, the threshold is also applied to the absolute value of negative values (generating negative spikes) |
|
|
Neural leak time constant \(\tau_{leak}\) (if 0, no leak) |
|
|
Neural refractory period \(T_{refrac}\) |
|
|
Terminate delta |
|
|
Relative initial synaptic weight \(w_{init}\) |
|
|
Mean minimum synaptic weight \(w_{min}\) |
|
|
Mean maximum synaptic weight \(w_{max}\) |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
OXRAM specific parameter |
|
|
Intrinsic SET switching probability \(P_{SET}\) (upon receiving a SET programming pulse). Assuming uniform statistical distribution (not well supported by experiments on RRAM) |
|
|
Intrinsic RESET switching probability \(P_{RESET}\) (upon receiving a RESET programming pulse). Assuming uniform statistical distribution (not well supported by experiments on RRAM) |
|
|
Synaptic redundancy (number of RRAM device per synapse) |
|
|
Bipolar weights |
|
|
Bipolar integration |
|
|
Extrinsic STDP LTP probability (cumulative with intrinsic SET switching probability \(P_{SET}\)) |
|
|
Extrinsic STDP LTD probability (cumulative with intrinsic RESET switching probability \(P_{RESET}\)) |
|
|
STDP LTP time window \(T_{LTP}\) |
|
|
Neural lateral inhibition period \(T_{inhibit}\) |
|
|
If false, STDP is disabled (no synaptic weight change) |
|
|
If true, reset the integration to 0 during the refractory period |
|
|
If false, the analog value of the devices is integrated, instead of their binary value |
LRN¶
Local Response Normalization (LRN) layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
The response-normalized activity \(b_{x,y}^{i}\) is given by the expression:
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Normalization window width in elements |
|
all Frame |
Value of the alpha variance scaling parameter in the normalization formula |
|
all Frame |
Value of the beta power parameter in the normalization formula |
|
all Frame |
Value of the k parameter in normalization formula |
LSTM¶
Long Short Term Memory Layer [HS97].
Global layer parameters (Frame_CUDA models)¶
Option [default value] |
Description |
|---|---|
|
Maximum sequence length that the LSTM can take as an input. |
|
Number of sequences used for a single weights actualisation process : size of the batch. |
|
Dimension of every element composing a sequence. |
|
Dimension of the LSTM inner state and output. |
|
If disabled return the full output sequence. |
|
If enabled, build a bidirectional structure. |
|
All Gates weights initial values filler. |
|
All Gates bias initial values filler. |
|
Input gate previous layer and recurrent weights initial values filler. Take precedence over AllGatesWeightsFiller parameter. |
|
Forget gate previous layer and recurrent weights initial values filler. Take precedence over AllGatesWeightsFiller parameter. |
|
Cell gate (or new memory) previous layer and recurrent weights initial values filler. Take precedence over AllGatesWeightsFiller parameter. |
|
Output gate previous layer and recurrent weights initial values filler. Take precedence over AllGatesWeightsFiller parameter. |
|
Input gate previous layer and recurrent bias initial values filler. Take precedence over AllGatesBiasFiller parameter. |
|
Forget gate recurrent bias initial values filler. Take precedence over AllGatesBiasFiller parameter. Often set to 1.0 to show better convergence performance. |
|
Forget gate previous layer bias initial values filler. Take precedence over AllGatesBiasFiller parameter. |
|
Cell gate (or new memory) previous layer and recurrent bias initial values filler. Take precedence over AllGatesBiasFiller parameter. |
|
Output gate previous layer and recurrent bias initial values filler. Take precedence over AllGatesBiasFiller parameter. |
|
Recurrent previous state initialisation. Often set to 0.0 |
|
Recurrent previous LSTM inner state initialisation. Often set to 0.0 |
Configuration parameters (Frame_CUDA models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Any solver parameters |
|
all Frame |
The probability with which the value from input would be dropped. |
|
all Frame |
If enabled, drop the matrix multiplication of the input data. |
|
all Frame |
Allow to choose different cuDNN implementation. Can be 0 : STANDARD, 1 : STATIC, 2 : DYNAMIC. Case 1 and 2 aren’t supported yet. |
Current restrictions¶
Only Frame_Cuda version is supported yet.
The implementation only support input sequences with a fixed length associated with a single label.
CuDNN structures requires the input data to be ordered as [1, InputDim, BatchSize, SeqLength]. Depending on the use case (like sequential-MNIST), the input data would need to be shuffled between the stimuli provisder and the RNN in order to process batches of data. No shuffling layer is yet operational. In that case, set batch to one for first experiments.
Further development requirements¶
When it comes to RNN, two main factors needs to be considered to build proper interfaces :
Whether the input data has a variable or a fixed length over the data base, that is to say whether the input data will have a variable or fixed Sequence length. Of course the main strength of a RNN is to process variable length data.
Labelling granularity of the input data, that is to say wheteher every elements of a sequence is labelled or the sequence itself has only one label.
For instance, let’s consider sentences as sequences of words in which every word would be part of a vocabulary. Sentences could have a variable length and every element/word would have a label. In that case, every relevant element of the output sequence from the recurrent structure is turned into a prediction throught a fully connected layer with a linear activation fonction and a softmax.
On the opposite, using sequential-MNIST database, the sequence length would be the same regarding every image and there is only one label for an image. In that case, the last element of the output sequence is the most relevant one to be turned into a prediction as it carries the information of the entire input sequence.
To provide flexibility according to these factors, the first implementation choice is to set a maximum sequence length emphSeqLength as an hyperparameter that the User provide. Variable length senquences can be processed by padding the remaining steps of the input sequence.
Then two cases occur as the labeling granularity is scaled at each element of the sequence or scaled at the sequence itself:
The sequence itself has only one label :
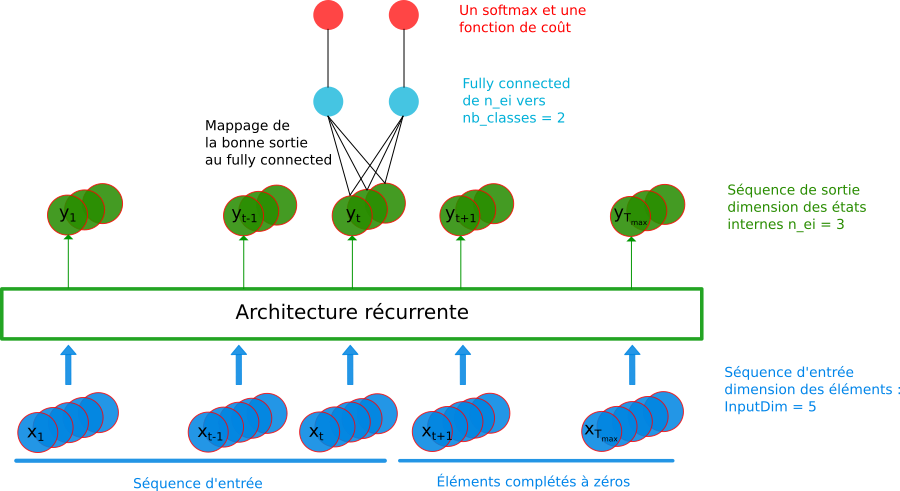
RNN model : variable sequence length and labeling scaled at the sequence¶
The model has a fixed size with one fully connected mapped to the relevant element of the output sequence according to the input sequence.Every elements of a sequence is labelled :
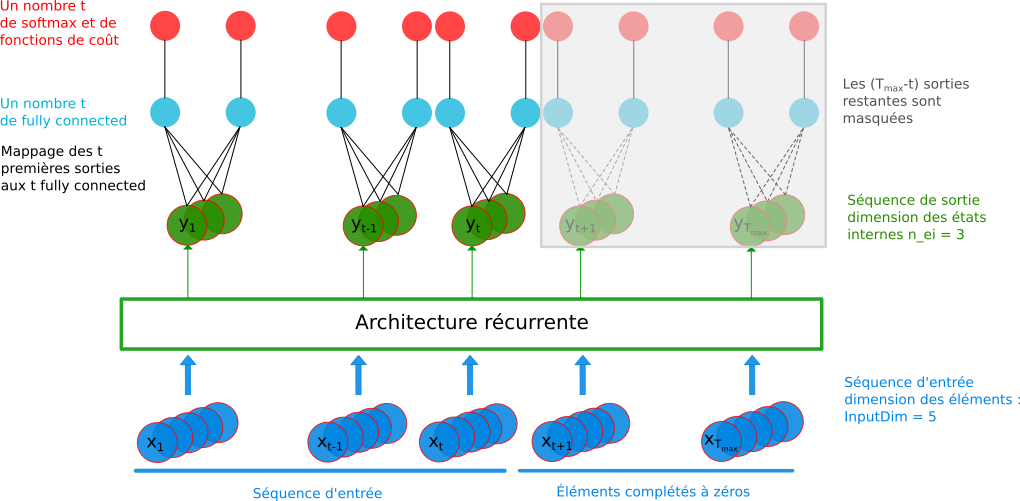
RNN model : variable sequence length and labeling scaled at each element of the sequence¶
Development guidance¶
Replace the inner local variables of LSTMCell_Frame_Cuda with a generic layer of shuffling (on device) to enable the the process of data batch.
Develop some kind of label embedding within the layer to better articulate the labeling granularity of the input data.
Adapt structures to support the STATIC and DYNAMIC algorithm of cuDNN functions.
Normalize¶
Normalize layer.
Option [default value] |
Description |
|---|---|
|
Number of output feature maps |
|
Norm to be used. Can be, |
Padding¶
Padding layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Size of the top padding (positive or negative) |
|
Size of the bottom padding (positive or negative) |
|
Size of the left padding (positive or negative) |
|
Size of the right padding (positive or negative) |
Pool¶
Pooling layer.
There are two CUDA models for this cell:
Frame_CUDA, which uses CuDNN as back-end and only supports
one-to-one input to output map connection;
Frame_EXT_CUDA, which uses custom CUDA kernels and allows arbitrary
connections between input and output maps (and can therefore be used to
implement Maxout or both Maxout and Pooling simultaneously).
Maxout example¶
In the following INI section, one implements a Maxout between each consecutive pair of 8 input maps:
[maxout_layer]
Input=...
Type=Pool
Model=Frame_EXT_CUDA
PoolWidth=1
PoolHeight=1
NbOutputs=4
Pooling=Max
Mapping.SizeY=2
Mapping.StrideY=2
The layer connectivity is the following:
# i n p u t m a p |
1 |
X |
|||
2 |
X |
||||
3 |
X |
||||
4 |
X |
||||
5 |
X |
||||
6 |
X |
||||
7 |
X |
||||
8 |
X |
||||
1 |
2 |
3 |
4 |
||
# output map |
|||||
Option [default value] |
Description |
|---|---|
|
Type of pooling ( |
|
Width of the pooling area |
|
Height of the pooling area |
|
Depth of the pooling area (implies 3D pooling area) |
|
Pooling area size (implies 2D square pooling area) |
|
List of space-separated dimensions for N-D pooling area |
|
Number of output channels |
|
X-axis stride of the pooling area |
|
Y-axis stride of the pooling area |
|
Z-axis stride of the pooling area |
|
Stride of the pooling area |
|
List of space-separated stride dimensions for N-D pooling area |
|
X-axis input padding |
|
Y-axis input padding |
|
Z-axis input padding |
|
Input padding |
|
List of space-separated padding dimensions for N-D pooling area |
|
Activation function. Can be any of |
|
Mapping: number of groups (mutually exclusive with all other Mapping.* options) |
|
Mapping: number of channels per group (mutually exclusive with all other Mapping.* options) |
|
Mapping canvas pattern default width |
|
Mapping canvas pattern default height |
|
Mapping canvas pattern default size (mutually
exclusive with |
|
Mapping canvas default X-axis step |
|
Mapping canvas default Y-axis step |
|
Mapping canvas default step (mutually exclusive
with``Mapping.StrideX`` and |
|
Mapping canvas default X-axis offset |
|
Mapping canvas default Y-axis offset |
|
Mapping canvas default offset (mutually exclusive
with |
|
Mapping canvas pattern default number of iterations (0 means no limit) |
|
Mapping canvas pattern default width for
input layer |
|
Mapping canvas pattern default height for
input layer |
|
Mapping canvas pattern default size for
input layer |
|
Mapping canvas default X-axis step for
input layer |
|
Mapping canvas default Y-axis step for
input layer |
|
Mapping canvas default step for input layer |
|
Mapping canvas default X-axis offset for
input layer |
|
Mapping canvas default Y-axis offset for
input layer |
|
Mapping canvas default offset for input
layer |
|
Mapping canvas pattern default number of
iterations for input layer |
Configuration parameters (Spike models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Spike |
Synaptic incoming delay \(w_{delay}\) |
value |
Rbf¶
Radial basis function fully connected layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Centers initial values filler |
[ |
|
|
Scaling initial values filler |
[ |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Any solver parameters |
|
all Frame |
Centers solver parameters, take precedence over the |
|
all Frame |
Scaling solver parameters, take precedence over the |
|
|
Approximation for the Gaussian function, can be any of: |
Resize¶
Resize layer can be applied to change dimension of features maps or of stimuli provider.
Option [default value] |
Description |
|---|---|
|
Number of output feature maps |
|
Output height dimension |
|
Output width dimension |
|
Resize interpolation mode. Can be, |
Configuration parameters¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Corner alignement mode if |
Softmax¶
Softmax layer.
Option [default value] |
Description |
|---|---|
|
Number of output neurons |
|
Softmax followed with a multinomial logistic layer |
|
Softmax is applied on groups of outputs. The group size must be a divisor of |
The softmax function performs the following operation, with \(a_{x,y}^{i}\) and \(b_{x,y}^{i}\) the input and the output respectively at position \((x,y)\) on channel \(i\):
and
When the WithLoss option is enabled, compute the gradient directly
in respect of the cross-entropy loss:
In this case, the gradient output becomes:
with
Transformation¶
Transformation layer, which can apply any transformation described in [sec:transformations]. Useful for fully CNN post-processing for example.
Option [default value] |
Description |
|---|---|
|
Number of outputs |
|
Name of the transformation to apply |
The Transformation options must be placed in the same section.
Usage example for fully CNNs:
[post.Transformation-thres]
Input=... ; for example, network's logistic of softmax output layer
NbOutputs=1
Type=Transformation
Transformation=ThresholdTransformation
Operation=ToZero
Threshold=0.75
[post.Transformation-morpho]
Input=post.Transformation-thres
NbOutputs=1
Type=Transformation
Transformation=MorphologyTransformation
Operation=Opening
Size=3
Threshold¶
Apply a thresholding.
Option [default value] |
Description |
|---|---|
|
Number of output feature maps |
|
Threshold value |
Configuration parameters (Frame models)¶
Option [default value] |
Model(s) |
Description |
|---|---|---|
|
all Frame |
Thresholding operation to apply. Can be: |
|
||
|
||
|
||
|
||
|
||
|
all Frame |
Max. value to use with |
Unpool¶
Unpooling layer.
Option [default value] |
Description |
|---|---|
|
Type of pooling ( |
|
Width of the pooling area |
|
Height of the pooling area |
|
Depth of the pooling area (implies 3D pooling area) |
|
Pooling area size (implies 2D square pooling area) |
|
List of space-separated dimensions for N-D pooling area |
|
Number of output channels |
|
Name of the associated pool layer for the argmax (the pool layer input and the unpool layer output dimension must match) |
|
X-axis stride of the pooling area |
|
Y-axis stride of the pooling area |
|
Z-axis stride of the pooling area |
|
Stride of the pooling area |
|
List of space-separated stride dimensions for N-D pooling area |
|
X-axis input padding |
|
Y-axis input padding |
|
Z-axis input padding |
|
Input padding |
|
List of space-separated padding dimensions for N-D pooling area |
|
Activation function. Can be any of |
|
Mapping: number of groups (mutually exclusive with all other Mapping.* options) |
|
Mapping: number of channels per group (mutually exclusive with all other Mapping.* options) |
|
Mapping canvas pattern default width |
|
Mapping canvas pattern default height |
|
Mapping canvas pattern default size (mutually
exclusive with |
|
Mapping canvas default X-axis step |
|
Mapping canvas default Y-axis step |
|
Mapping canvas default step (mutually exclusive
with``Mapping.StrideX`` and |
|
Mapping canvas default X-axis offset |
|
Mapping canvas default Y-axis offset |
|
Mapping canvas default offset (mutually exclusive
with |
|
Mapping canvas pattern default number of iterations (0 means no limit) |
|
Mapping canvas pattern default width for
input layer |
|
Mapping canvas pattern default height for
input layer |
|
Mapping canvas pattern default size for
input layer |
|
Mapping canvas default X-axis step for
input layer |
|
Mapping canvas default Y-axis step for
input layer |
|
Mapping canvas default step for input layer |
|
Mapping canvas default X-axis offset for
input layer |
|
Mapping canvas default Y-axis offset for
input layer |
|
Mapping canvas default offset for input
layer |
|
Mapping canvas pattern default number of
iterations for input layer |